# Generate distributions with different kurtosis
set.seed(123)
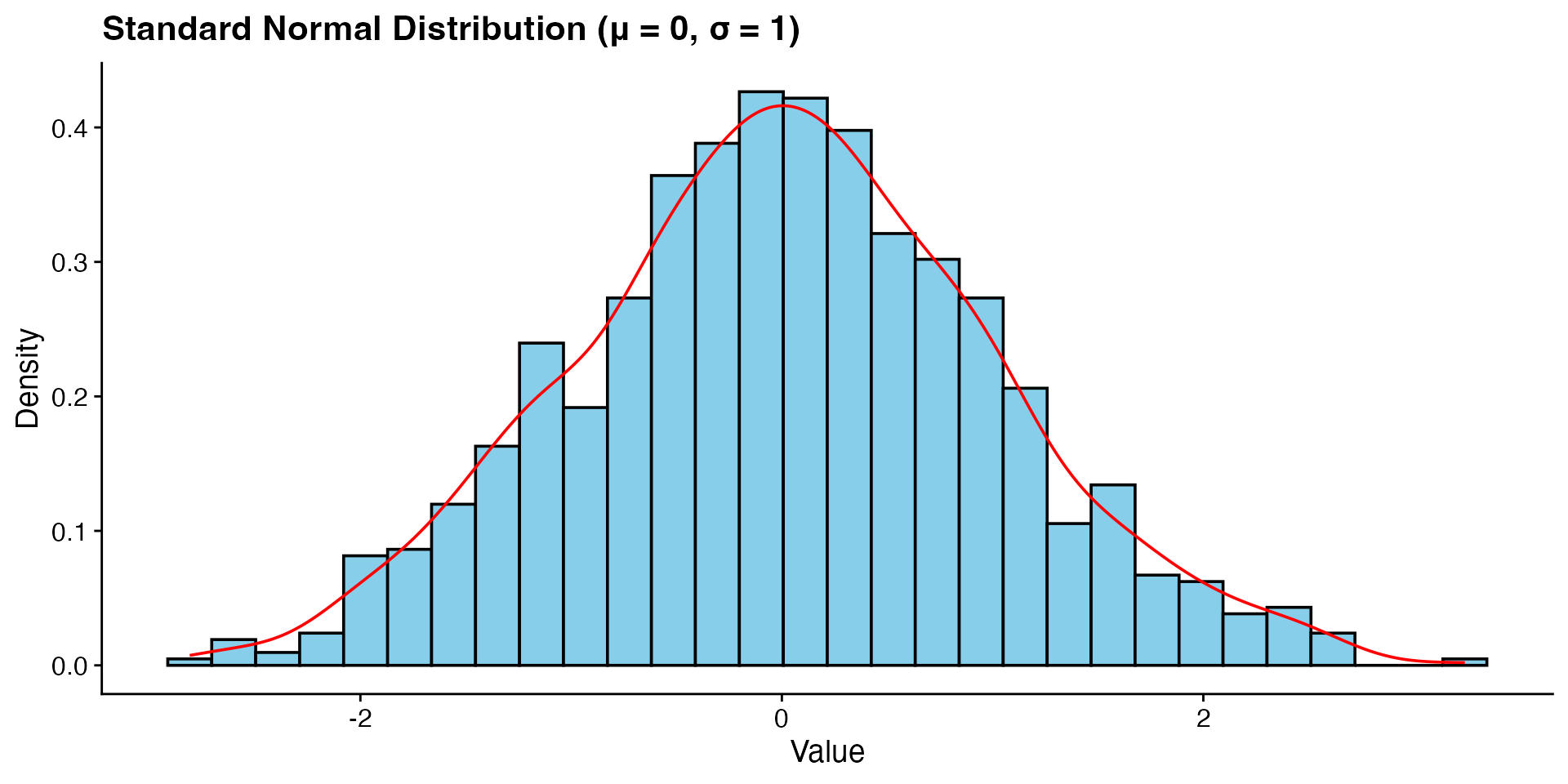
normal <- rnorm(1000, 0, 1) # Mesokurtic
leptokurtic <- rt(1000, df = 5) # t-distribution with 5 df is leptokurtic
platykurtic <- runif(1000, -3, 3) # Uniform distribution is platykurtic
# Calculate kurtosis values (using e1071 package)
library(e1071)
k_normal <- kurtosis(normal)
k_lepto <- kurtosis(leptokurtic)
k_platy <- kurtosis(platykurtic)
# Plot distributions
p1 <- ggplot(data.frame(x = normal), aes(x = x)) +
geom_histogram(aes(y = after_stat(density)), bins = 30,
fill = "#a6cee3", # colourblind-friendly blue
colour = "black") +
geom_density(colour = "#1f78b4", linewidth = 1) + # Darker blue
annotate("text", x = 2, y = 0.3,
label = paste("Kurtosis =", round(k_normal, 2)),
colour = "#1f78b4") +
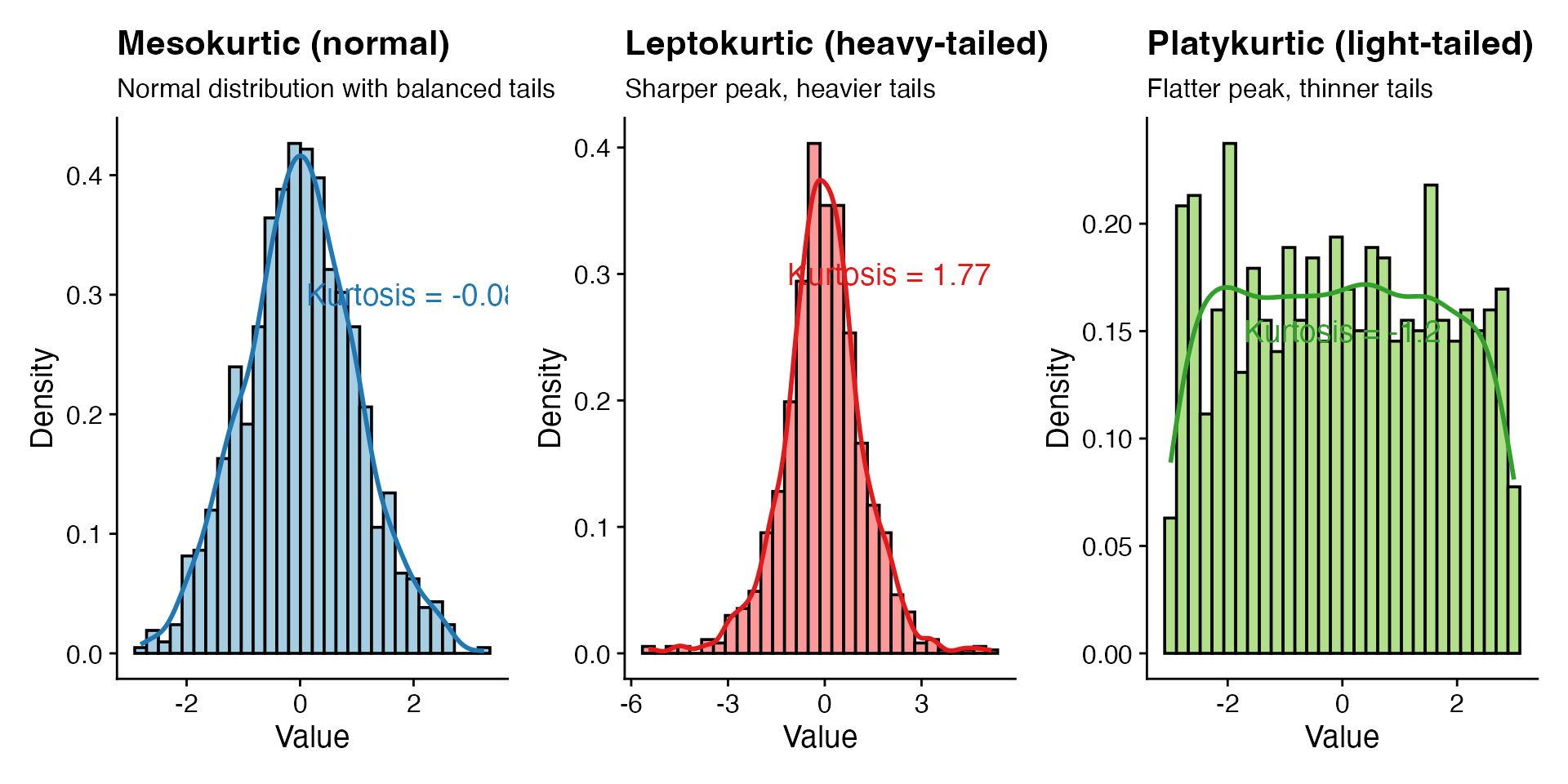
labs(title = "Mesokurtic (normal)",
subtitle = "Normal distribution with balanced tails",
x = "Value",
y = "Density")
p2 <- ggplot(data.frame(x = leptokurtic), aes(x = x)) +
geom_histogram(aes(y = after_stat(density)), bins = 30,
fill = "#fb9a99", # colourblind-friendly pink
colour = "black") +
geom_density(colour = "#e31a1c", linewidth = 1) + # Darker red
annotate("text", x = 2, y = 0.3,
label = paste("Kurtosis =", round(k_lepto, 2)),
colour = "#e31a1c") +
labs(title = "Leptokurtic (heavy-tailed)",
subtitle = "Sharper peak, heavier tails",
x = "Value",
y = "Density")
p3 <- ggplot(data.frame(x = platykurtic), aes(x = x)) +
geom_histogram(aes(y = after_stat(density)), bins = 30,
fill = "#b2df8a", # colourblind-friendly green
colour = "black") +
geom_density(colour = "#33a02c", linewidth = 1) + # Darker green
annotate("text", x = 0, y = 0.15,
label = paste("Kurtosis =", round(k_platy, 2)),
colour = "#33a02c") +
labs(title = "Platykurtic (light-tailed)",
subtitle = "Flatter peak, thinner tails",
x = "Value",
y = "Density")
# Display plots vertically
library(patchwork)
p1 + p2 + p3