penguins <- palmerpenguins::penguins
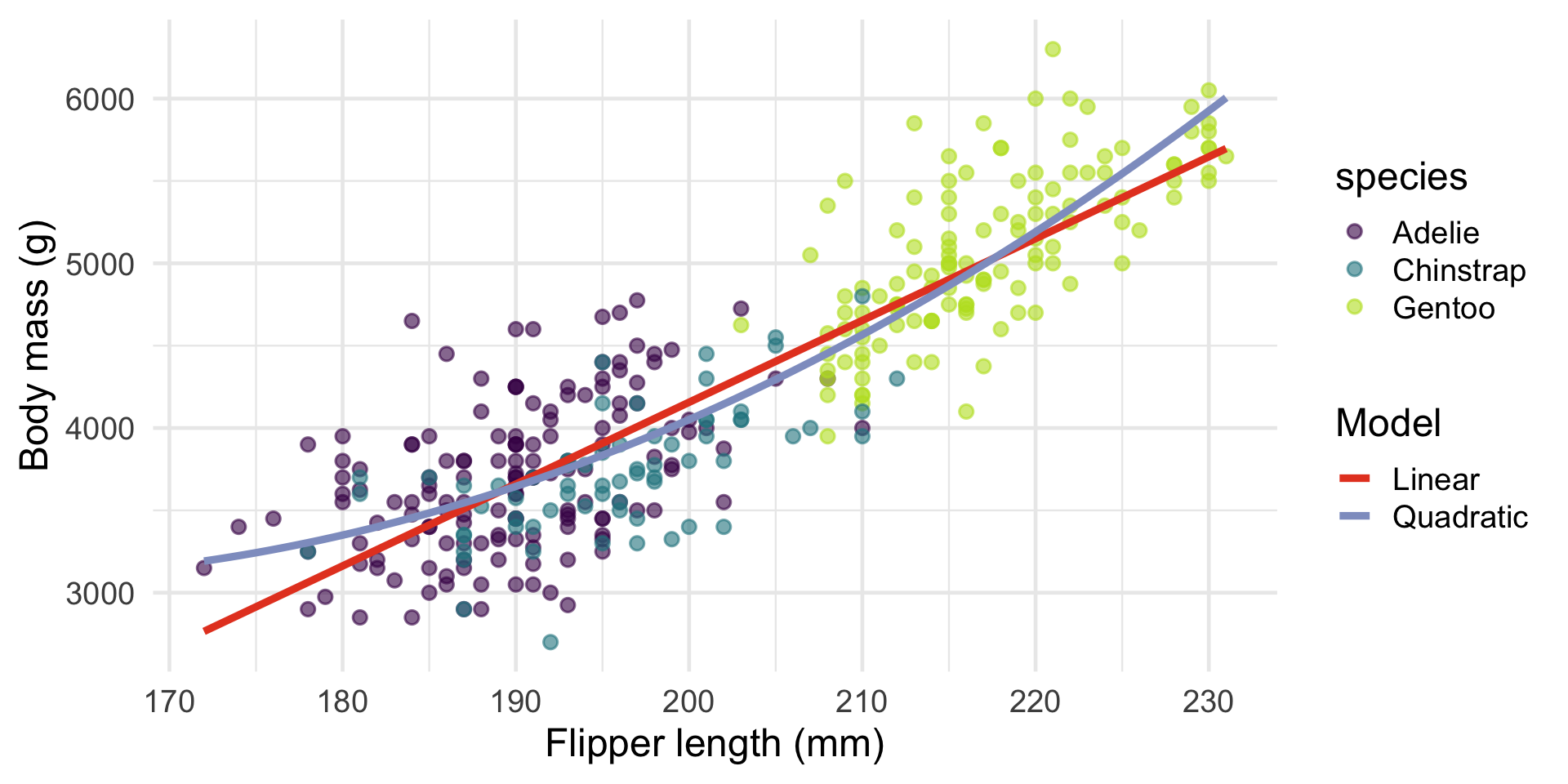
ggplot(penguins, aes(x = flipper_length_mm, y = body_mass_g)) +
geom_point(aes(colour = species), alpha = 0.6) +
geom_smooth(method = "lm", se = FALSE, colour = "#e64626", aes(linetype = "Linear")) +
geom_smooth(method = "lm", formula = y ~ poly(x, 2), se = FALSE, colour = "#8f9ec9", aes(linetype = "Quadratic")) +
scale_colour_viridis_d(end = 0.9) +
scale_linetype_manual(values = c(Linear = "solid", Quadratic = "solid")) +
guides(linetype = guide_legend(override.aes = list(colour = c("#e64626", "#8f9ec9")))) +
labs(x = "Flipper length (mm)", y = "Body mass (g)", linetype = "Model") +
theme_minimal(base_size = 18)