Lecture 03a – \(t\)-Tests
ENVX2001 Applied Statistical Methods
Apr 2026
The \(t\)-test (revision)
Introduction
- One-sample \(t\)-test: compare a sample mean against a known value
- Two-sample \(t\)-test: compare the means of two independent groups
- Paired \(t\)-test: compare measurements taken on the same subjects (e.g. before and after)
Last week we learned about confidence intervals, which estimate the uncertainty of a parameter (e.g. a mean).
The \(t\)-test and the confidence interval are closely related. Both are derived from the same quantities, and this week we look at that connection more carefully.
Cattle
Two breeds of cattle, 12 and 15 samples. Are the breeds different in mean weight?
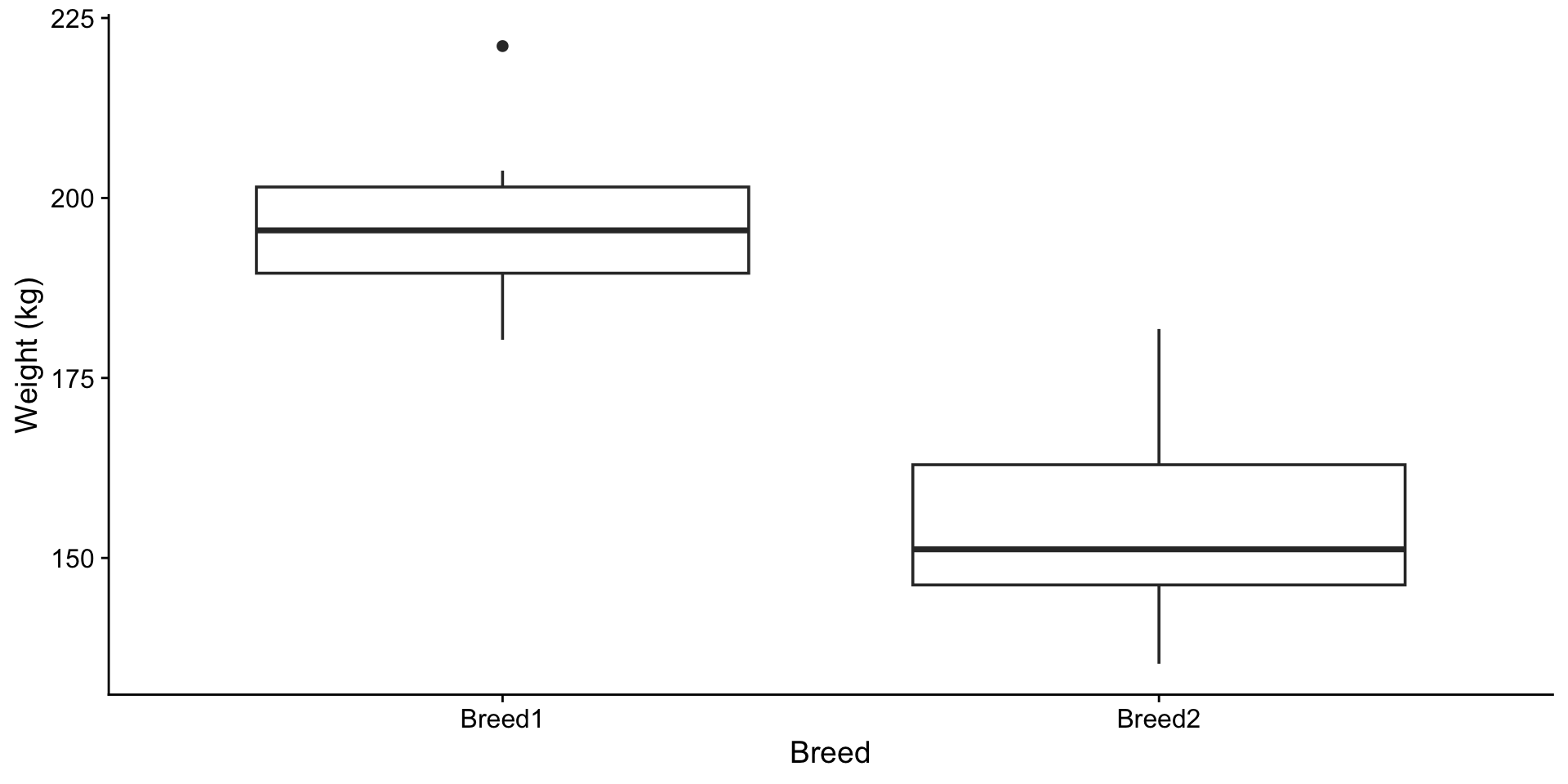
The boxplot suggests a difference. To test, you would run t.test() and check the p-value. What is the test actually computing?
From CIs to the \(t\)-test
The difference between two means
In the last lecture, we constructed confidence intervals for a single mean. The same principle extends to the difference between two means.
- Estimate the difference: \(\bar{y}_1 - \bar{y}_2\)
- Construct a 95% confidence interval around that estimate
- Evaluate whether the interval includes zero
- If the interval excludes zero: the means are significantly different at the 5% level
- If the interval includes zero: the data are consistent with no difference
t.test() gives you both
Two Sample t-test
data: weight by breed
t = 9.4624, df = 25, p-value = 9.663e-10
alternative hypothesis: true difference in means between group Breed1 and group Breed2 is not equal to 0
95 percent confidence interval:
33.23011 51.71989
sample estimates:
mean in group Breed1 mean in group Breed2
196.175 153.700 The output reports two quantities:
- A p-value (probability of observing a result this extreme if the null hypothesis were true)
- A 95% confidence interval for the difference between the two means
These always agree. If the CI excludes zero, the p-value is below 0.05, and vice versa. The \(t\)-statistic measures how far the observed difference is from zero, relative to its standard error.
Exploring the confidence interval
- Increasing the difference between means moves the CI away from zero
- Decreasing variability narrows the CI
- When the interval excludes zero, the means are significantly different
The CI for a difference
In the last lecture, we built a 95% CI for a single mean:
\[\bar{y} \pm t^* \times SE\]
For the difference between two means, the same structure applies:
\[(\bar{y}_1 - \bar{y}_2) \pm t^* \times SE(\bar{y}_1 - \bar{y}_2)\]
If this interval excludes zero, the groups differ significantly.
The \(t\)-statistic
We can rearrange the CI formula to isolate a single number that captures both the difference and the variability:
\[t = \frac{\bar{y}_1 - \bar{y}_2}{SE(\bar{y}_1 - \bar{y}_2)}\]
The \(t\)-statistic counts how many standard errors the observed difference is from zero.
For the cattle data: \(t = 42.5 / 4.5 = 9.46\). The observed difference is over nine standard errors from zero.
From the \(t\)-statistic to the p-value
We have a \(t\)-statistic. The p-value answers: if there were truly no difference between the breeds, how often would we see a value of \(|t|\) this large or larger?
- A small p-value means the observed difference is hard to explain by chance alone
- A large p-value means the observed difference is consistent with random variation
The 95% CI and the p-value always agree. If the 95% CI excludes zero, the p-value is below 0.05, and vice versa.
Exploring the p-value
- The shaded area under both tails is the p-value
- As the \(t\)-statistic moves further from zero, the p-value decreases
- At low degrees of freedom the tails are heavier, but the shape approaches the normal as df increases
The hypothesis testing workflow
Step 1: State what we are testing
- Null hypothesis: \(H_0: \mu_1 = \mu_2\) (no difference in mean weight between breeds)
- Alternative hypothesis: \(H_1: \mu_1 \neq \mu_2\) (means differ)
- This is a two-tailed test: we are looking for a difference in either direction (Breed 1 heavier or lighter than Breed 2), not just one
Model equation: \(y_{ij} = \mu_i + \varepsilon_{ij}\)
Each observation is the group mean plus random error. The hypothesis asks whether \(\mu_1\) and \(\mu_2\) are equal.
Step 2a: Equal variances
The \(t\)-test assumes both groups have similar variability. If they do not, then the test is unreliable because the pooled estimate of variance will not represent either group well.
- Rule of thumb: the ratio of the larger SD to the smaller SD should be below 2.0
Code
# A tibble: 2 × 4
breed n mean_wt sd_wt
<fct> <int> <dbl> <dbl>
1 Breed1 12 196. 10.6
2 Breed2 15 154. 12.3[1] 1.158756Step 2b: Normality
The \(t\)-test assumes the data in each group follow a normal distribution. With small samples, departures from normality (strong skew, heavy tails, outliers) can distort the p-value and confidence interval. Check each group separately.
Shapiro-Wilk normality test
data: cattle$weight[cattle$breed == "Breed1"]
W = 0.94084, p-value = 0.5091
Shapiro-Wilk normality test
data: cattle$weight[cattle$breed == "Breed2"]
W = 0.94636, p-value = 0.4691Step 2b: Normality (QQ plots)
Code
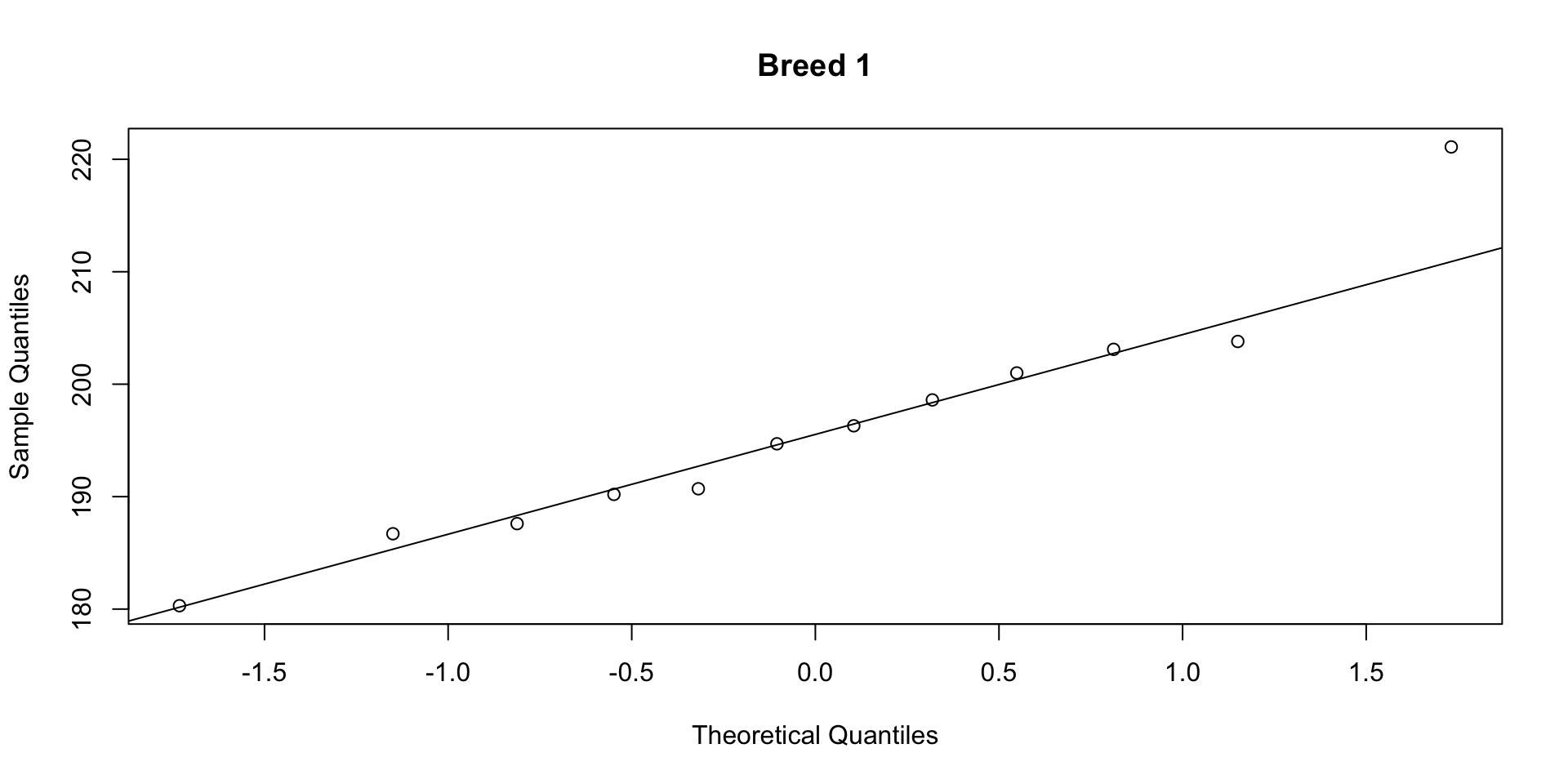
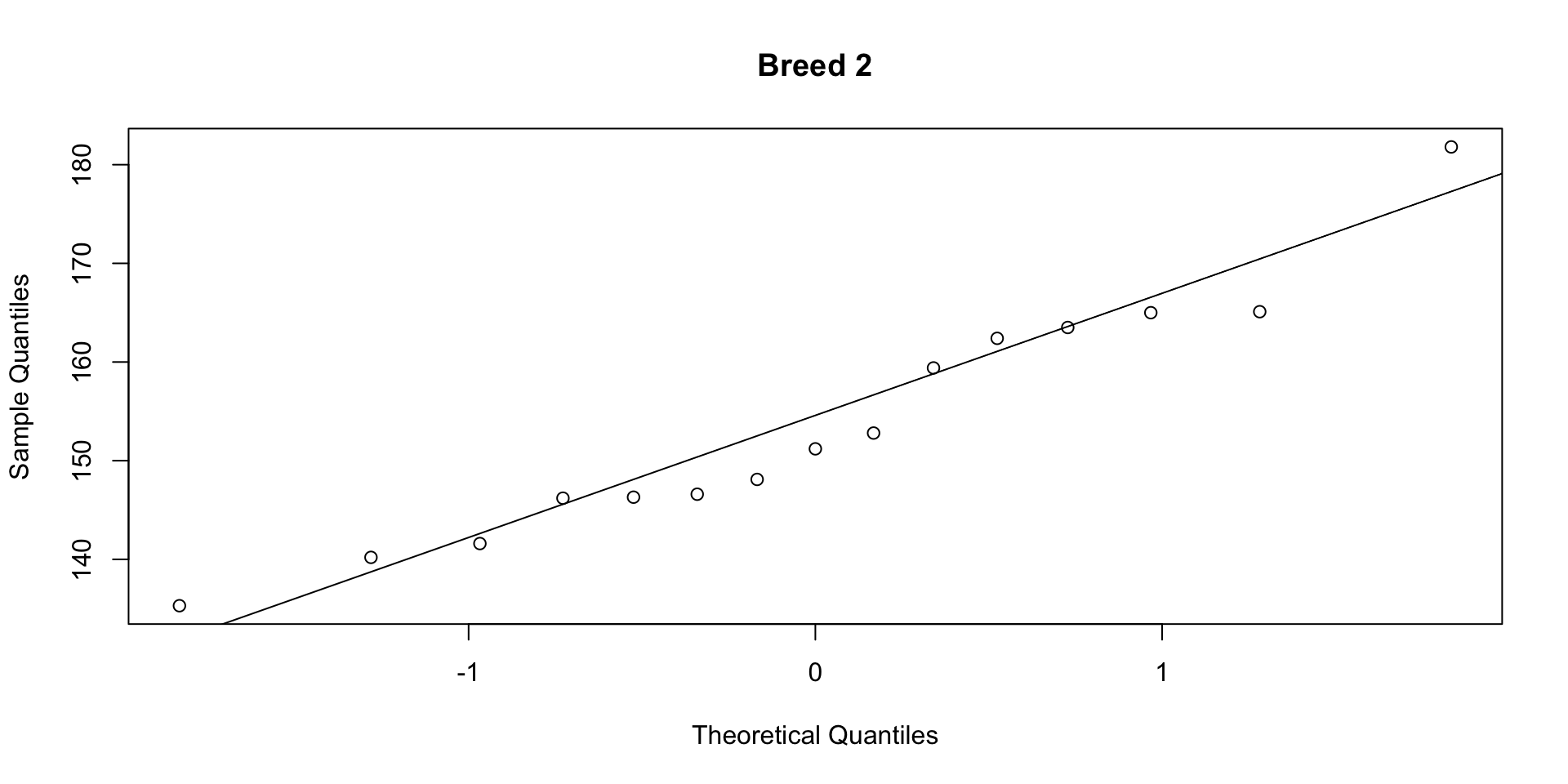
If points fall close to the line, the data are approximately normal. Both groups pass the Shapiro-Wilk test (\(p > 0.05\)) and the QQ plots show no strong departures.
Step 2c: Independence
Observations should not influence each other. This is a property of the study design, not something we test statistically.
- Each animal is measured once
- The weight of one animal does not affect the weight of another
- If the same animals were measured before and after a treatment, observations would not be independent (use a paired \(t\)-test instead)
Step 3: Run the test and interpret
Two Sample t-test
data: weight by breed
t = 9.4624, df = 25, p-value = 9.663e-10
alternative hypothesis: true difference in means between group Breed1 and group Breed2 is not equal to 0
95 percent confidence interval:
33.23011 51.71989
sample estimates:
mean in group Breed1 mean in group Breed2
196.175 153.700 - The \(t\)-statistic, degrees of freedom (\(n_1 + n_2 - 2\)), and p-value are reported together
var.equal = TRUEperforms the pooled (Student’s) \(t\)-test, which assumes equal variances
A p-value of 0.03 does not mean there is a 3% chance the null hypothesis is true. It means: if there were truly no difference, we would observe a result this extreme about 3% of the time.
HATPC
Notice the five steps we just followed:
- Hypotheses: state the null and alternative
- Assumptions: check equal variances, normality, independence
- Test: run the appropriate test
- P-value: interpret the result
- Conclusion: state the finding in context
We will use this framework throughout the semester, starting with ANOVA in the next lecture.
What if assumptions are not met?
- Unequal variances: use Welch’s \(t\)-test (the default in R when
var.equal = FALSE) - Non-normality: if \(N > 30\), the Central Limit Theorem provides reasonable coverage
- Non-independence: consider a paired \(t\)-test (e.g. before and after measurements on the same subjects)
Summary
Key points
- The two-sample \(t\)-test compares means of two independent groups
- The \(t\)-test and the CI for the difference are two views of the same calculation
- Check the three assumptions (equal variances, normality, independence) before running any test
- Follow HATPC as a structured process for hypothesis testing
In Lab 03, you will practise this workflow with t.test() and diagnostic plots.
Next: what if we have more than two groups? We need ANOVA.
15 minute break
This presentation is based on the SOLES Quarto reveal.js template and is licensed under a Creative Commons Attribution 4.0 International License.